News
Members
Publications
Software / Data
Job offers
Images / Videos
Collaborations
Conferences
Lab meetings: "Les partages de midi"
Practical information
Members Area
Next conferences we are in …



This shows you the differences between two versions of the page.
| Both sides previous revision Previous revision Next revision | Previous revision Last revision Both sides next revision | ||
|
software [2020/10/22 15:01] ahuaulme |
software [2022/11/28 14:02] ahuaulme |
||
|---|---|---|---|
| Line 1: | Line 1: | ||
| <html><div class="pageTitle"> Available data-sets </div></html> | <html><div class="pageTitle"> Available data-sets </div></html> | ||
| + | |||
| + | ====== PETRAW ====== | ||
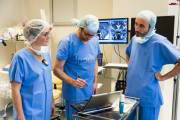
| + | The “PEg TRAnsfer Workflow recognition by different modalities” (PETRAW) data-set has been present as a sub-challenge of [[https://endovis.grand-challenge.org/|Endovis2021]] at [[https://www.miccai2021.org/en/|MICCAI2021]]. | ||
| + | |||
| + | |||
| + | The data-set contains 125 peg transfer sequences performed on a virtual simulator. Each sequence is composed of the following information: | ||
| + | * Video | ||
| + | * Kinematic data | ||
| + | * Segmentation | ||
| + | * Workflow annotation at 3 levels of granularity (phases, steps, and activities) | ||
| + | |||
| + | {{ ::petraw.jpg?nolink&400 |}} | ||
| + | |||
| + | You can found additional information and download the data-set here: https://www.synapse.org/MISAW | ||
| + | |||
| + | ====== MISAW ====== | ||
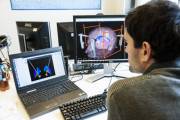
| + | The “MIcro-Surgical Anastomose Workflow recognition on training sessions” (MISAW) data-set has been present as a sub-challenge of [[https://endovis.grand-challenge.org/|Endovis2020]] at [[https://www.miccai2020.org/en/|MICCAI2020]]. | ||
| + | |||
| + | |||
| + | The data-set contains 27 micro-anastomosis training sequences and is composed of the following information: {{ :snapshots_micro_surgical_anastomose.png?200|MISAW data-set snapshots}} | ||
| + | * Stereoscopic video | ||
| + | * Kinematic data | ||
| + | * Workflow annotation at 3 levels of granularity (phases, steps, and activities) | ||
| + | |||
| + | You can found additional information and download the data-set here: https://www.synapse.org/MISAW | ||
| + | |||
| + | |||
| + | |||
| ====== Ontologies ====== | ====== Ontologies ====== | ||
| Line 8: | Line 36: | ||
| You can access the dataset or download directly corresponding images and annotations files [[https://ged.univ-rennes1.fr/nuxeo/nxpath/default/default-domain/workspaces/recherche/LTSI/MediCIS/NeuroSurgicalToolsDatase@view_documents|here]]. | You can access the dataset or download directly corresponding images and annotations files [[https://ged.univ-rennes1.fr/nuxeo/nxpath/default/default-domain/workspaces/recherche/LTSI/MediCIS/NeuroSurgicalToolsDatase@view_documents|here]]. | ||
| - | [[https://dbouget.bitbucket.io/2015_tmi_surgical_tool_detection/|{{ :dataneurosurg.png?300}}]] | + | [[https://dbouget.bitbucket.io/2015_tmi_surgical_tool_detection/|{{ :dataneurosurg.png?200}}]] |
| ====== MR mono-subject template ====== | ====== MR mono-subject template ====== | ||


